Scientists at the CSIR-Centre for Cellular and Molecular Biology (CCMB) in Hyderabad have analysed more than 2,000 SARS-CoV-2, the virus that causes COVID-19 genomes from India available in the public domain to understand the various strains in circulation.
Earlier in June, the team had revealed the presence of a distinct virus population among Indians.
This was named the clade I/A3i, and is recognised by the presence of 4 specific variations in their genetic makeup (genomes).
At that time, 41 per cent of all Indian SARS-CoV-2 genomes belonged to this clade.
However, the current analysis showed that the proportion of the A3i clade dropped to 18 per cent, it said.
The decrease in the proportion of A3i clade is accompanied by an increase of the A2a clade, also referred to as the G clade or the 20A/B/C clades in other nomenclatures.
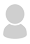
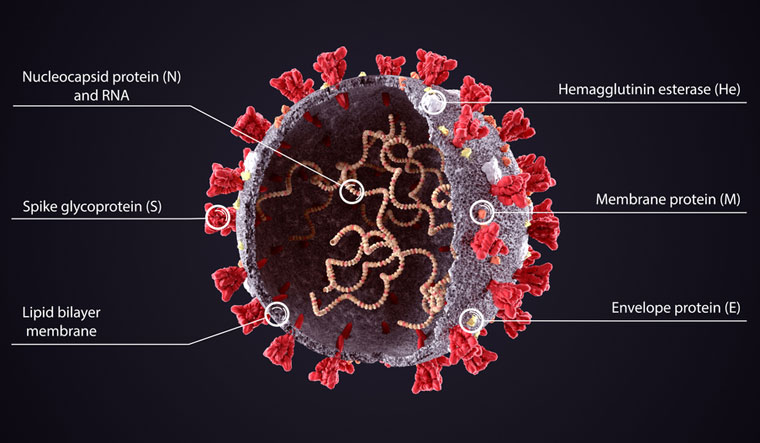
Viruses of the A2a or the G clade carry the D614G mutation in their spike protein which is shown to be associated with an increased infectivity.
At present, approximately 70 per cent of all Indian as well as global SARS-CoV-2 genomes fall into this clade.
"As expected for a strain which is more infectious, A2a clade quickly became the dominant clade in India just like everywhere else. There is no evidence to state that this mutation is clinically a more difficult one," Dr Rakesh K Mishra, Director, CCMB and a co-author of the study said.
"The similarity in viral genome globally should be considered a positive news, because a vaccine or a drug targeting this mutation will work with the same effectiveness all over the world," he added.
It is, however, important to note that no clade at present has been conclusively shown to be associated with a more severe form of COVID-9, or an increased risk of death.
The findings of the study done with scientists from CSIR-Institute of Integrative Biology as collaborators, are now peer-reviewed and published in the journal Open Forum Infectious Diseases published by the Oxford University Press, the CCMB release said.